Alex Price, Ph.D.
-
Assistant Professor, Genome Regulation and Cell Signaling Program, Ellen and Ronald Caplan Cancer Center
Price’s research focus is on how DNA viruses co-opt and manipulate cellular RNA processing pathways.
Growing up on the west coast, Price obtained his Bachelor’s of Science degree in Genetics and Cell Biology at Washington State University. Moving to the east coast, he obtained a Ph.D. in Molecular Genetics and Microbiology from Duke University in 2016. To pursue postdoctoral research, Price moved to Philadelphia, where he was associated with the University of Pennsylvania and the Children’s Hospital of Philadelphia. In 2023 he joined the Wistar Institute as an Assistant Professor in the Gene Expression and Regulation Program, which became the Genome Regulation and Cell Signaling Program in 2024.
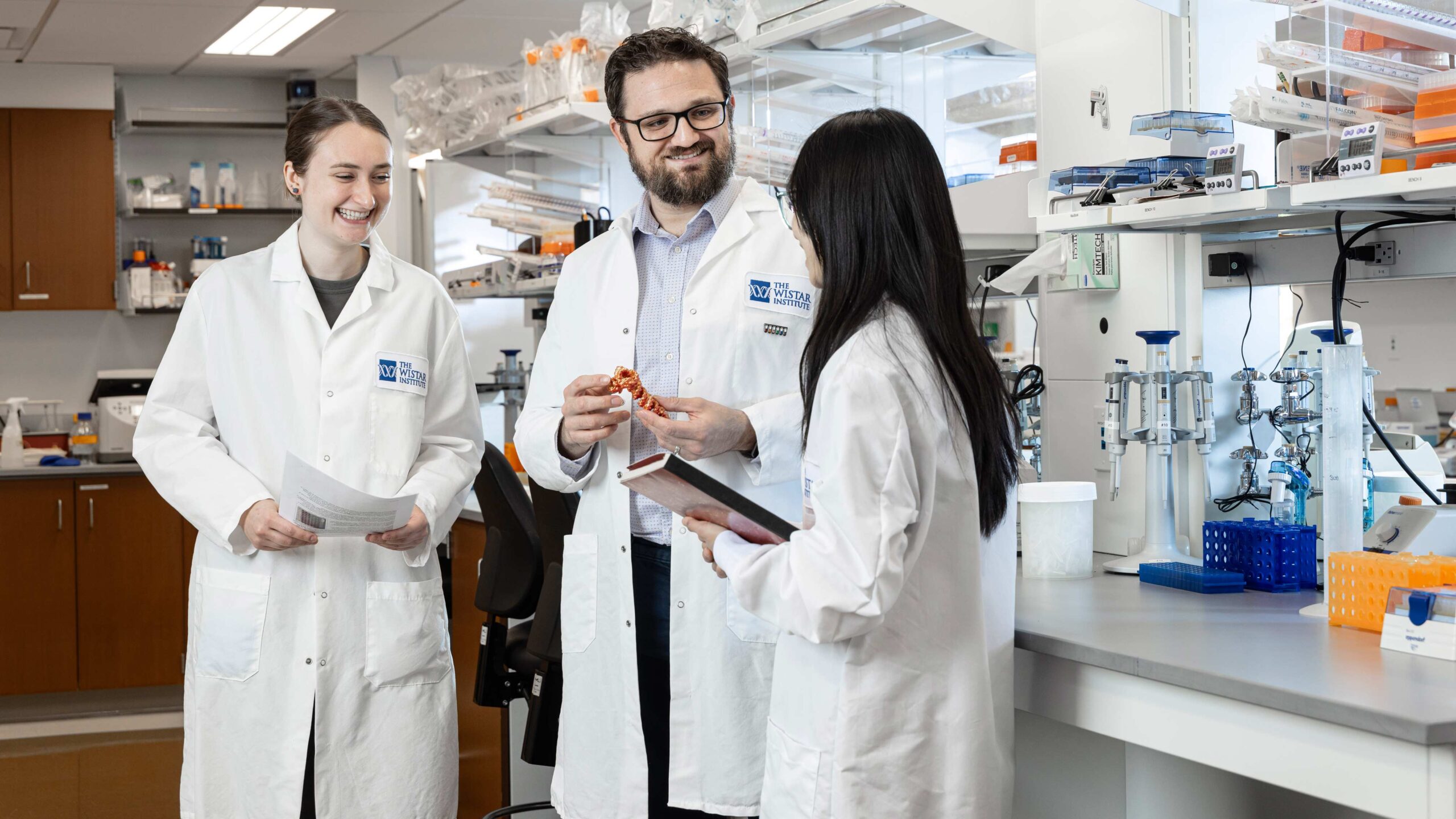
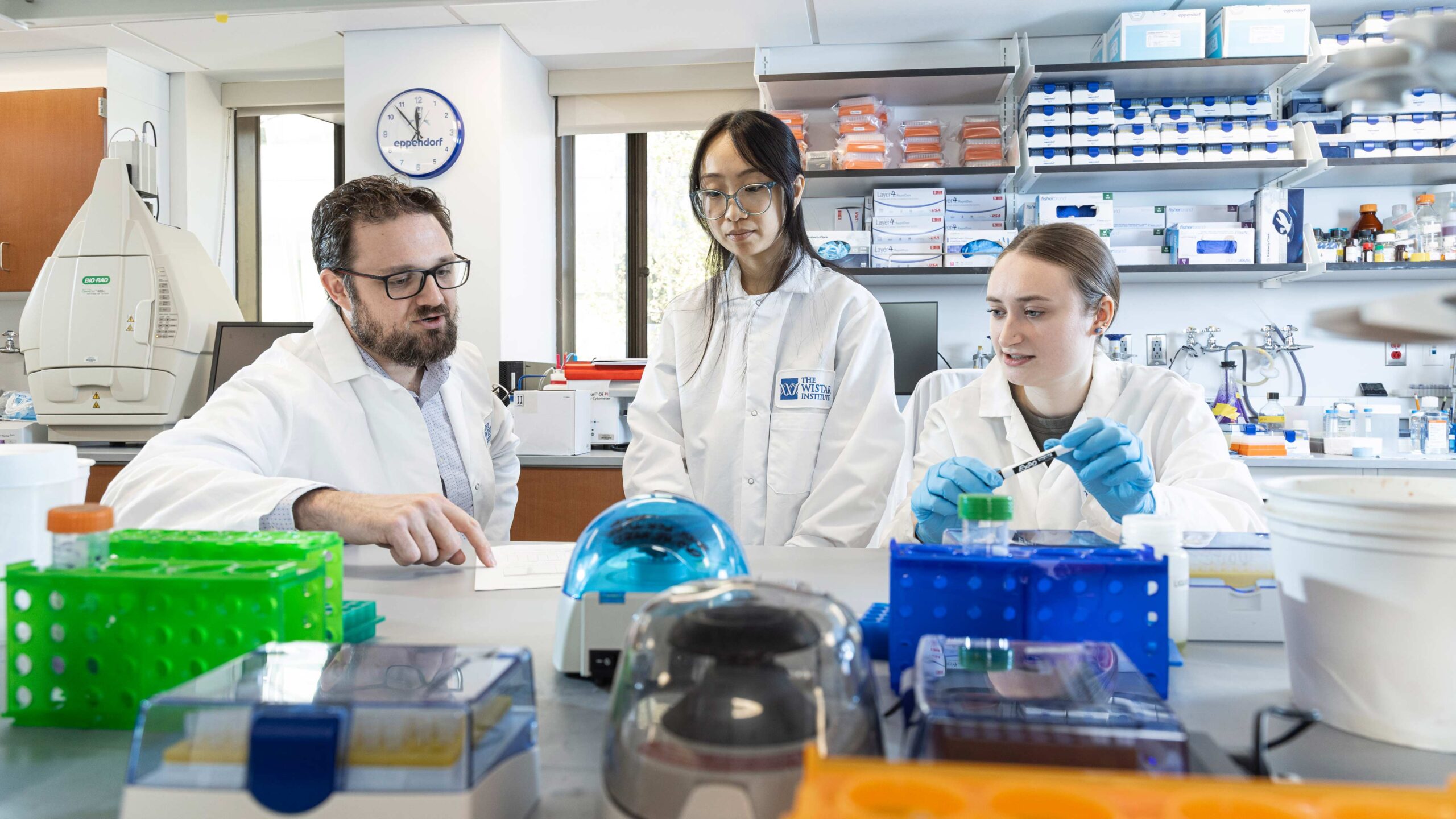
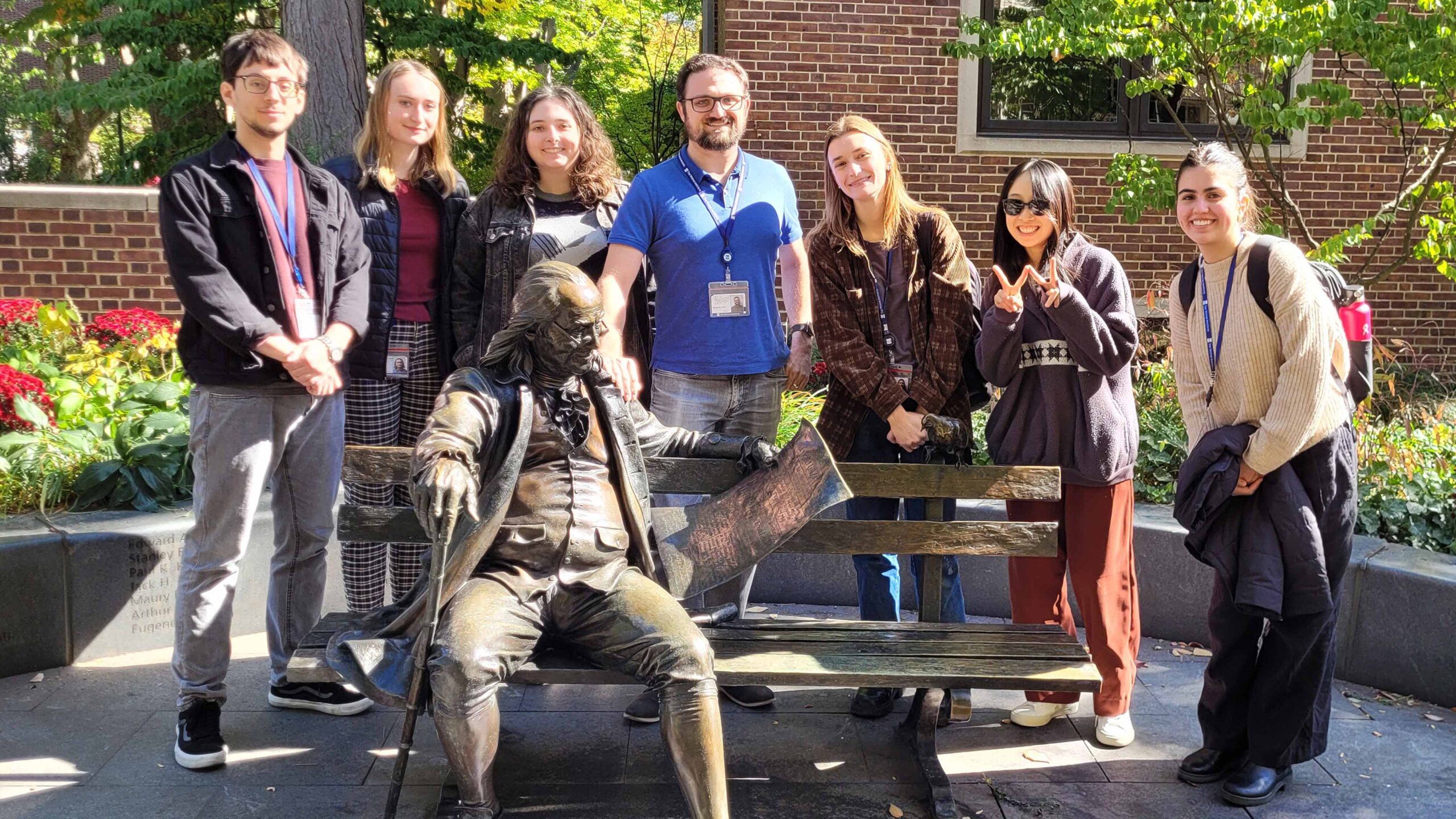
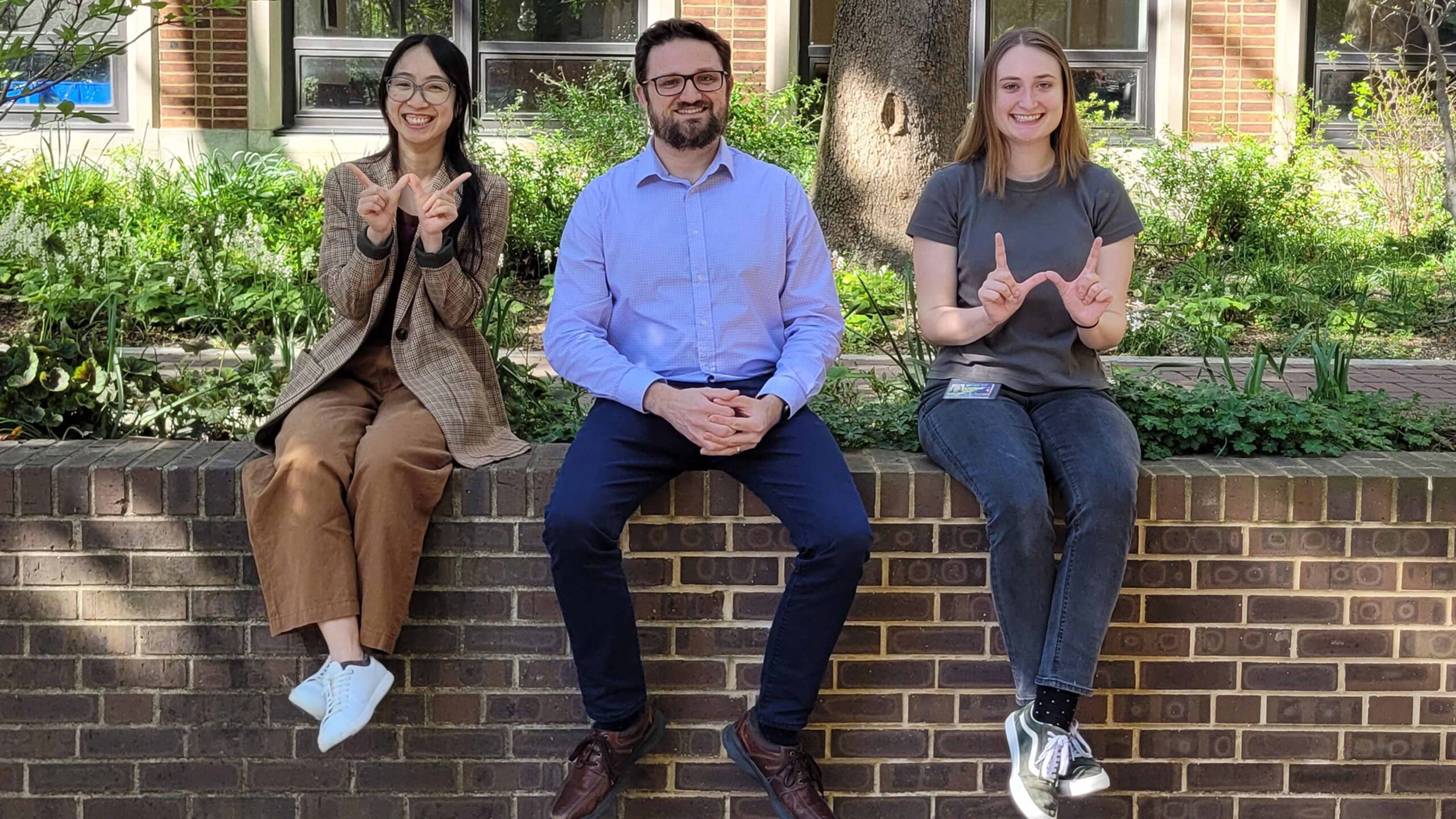
The Price Laboratory

The Price Laboratory
The Price Lab studies how DNA viruses take over and subvert host cell biology, shining light on the fundamental processes they must steal from their host to replicate. While viruses have evolved to be masters of molecular mimicry, any viral process that deviates from standard cell biology allows a host cell to sense infection. DNA viruses obligately use cellular RNA processing machinery to make viral transcripts, yet are constrained by a hard limit on maximum genome size. This means that viruses often express tens to hundreds of messages from coding space the size of a single cellular gene. By pushing cellular machinery to the absolute limit, these pathogens have adopted a high risk, high reward strategy for maximizing gene expression that can inadvertently activate the innate immune system.
Our goal is to discover how viruses balance the ability to produce diverse RNAs from limited coding capacity while preventing the deleterious formation of non-self RNAs. In doing so, we aim to uncover therapeutic vulnerabilities in the transcriptional programs enacted by diverse viruses. Beyond virology, our research will reveal how transcriptional processes affect innate immunity and inflammation, with broad implications for the progression of autoimmune diseases and cancers where RNA biogenesis has become dysregulated.
-
Postdoctoral Fellows
Alison Yu, Ph.D.
-
Graduate Students
Lorenzo Serra (UniBO)
-
Research Assistant
Claire O’Brien
-
Learn about job opportunities at The Wistar Institute here.
Research
Question 1: How is nuclear dsRNA sensed and responded to during viral infection?
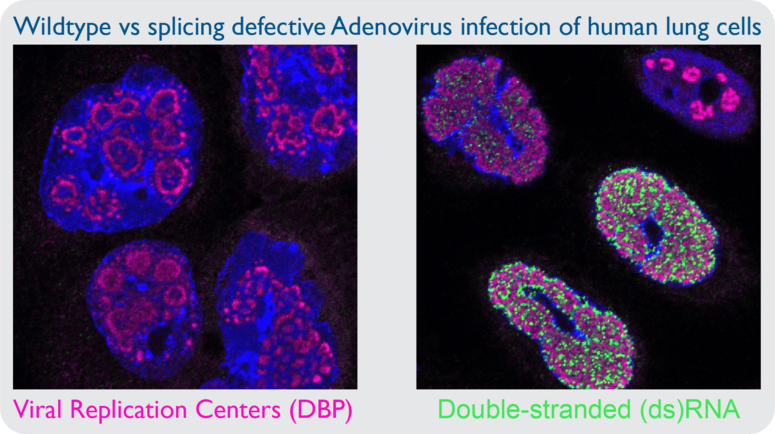
Viruses with limited genome size maximize coding capacity via regulated expression of genes on both DNA strands. It is thought that convergent transcription of DNA virus genomes leads to the production of dsRNA, and this postulate is supported by the fact that these viruses encode inhibitors of dsRNA sensing pathways. However, there is little primary evidence of dsRNA accumulating during DNA virus infection. In my prior work I utilized monoclonal antibodies directed against dsRNA to examine cells infected by adenovirus, yet I was unable to detect dsRNA. However, infection with adenovirus mutant viruses that exhibit inefficient splicing of viral transcripts leads to robust dsRNA production in the nucleus. The presence of nuclear dsRNA during mutant adenovirus infection is accompanied by activation of cytoplasmic sensors PKR and RNase L, yet there is no known nuclear sensor of dsRNA. Assessing the role of nuclear dsRNA sensing is becoming increasingly important as it is more appreciated that hyperactivity of cellular antiviral sensors can lead to autoimmune disease. Specific questions include:
- How Is nuclear dsRNA sensed by the innate immune system?
- How does nuclear recognition of dsRNA communicate with existing components of the immune system?
- How do nuclear dsRNA-binding proteins interact with viral dsRNA?
Question 2: How do DNA viruses spatially regulate RNA transcription and processing?
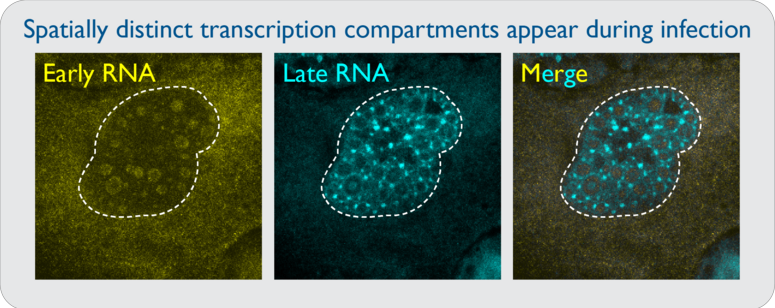
Unprocessed adenoviral RNAs are likely to form dsRNA if they are synthesized in close proximity to antisense transcripts. Viruses that can regulate sense and antisense transcription in spatially distinct nuclear compartments would allow for localized processing that would preclude dsRNA formation. It was previously shown that viral DNA replication takes place inside biophysical viral replication centers, and that RNA synthesis occurs in a shell surrounding these regions. However, how this spatial segregation occurs is unknown. Specific questions include:
- How are viral genomes spatially organized?
- How do DNA viruses spatially regulate RNA transcription?
- Does transcription of viral genomes lead to viral mutation or genome instability?
Question 3: How do herpesviruses exploit RNA processing?
Herpesvirus lytic transcripts are often intron-less and bypass many canonical processing steps on the path to expression. Viral RNA-binding proteins, whose functions are only recently being fully understood, often mediate these processes. In contrast, latent phase infection encodes complex transcripts with extensive alternative splicing reliant on cellular factors for expression. Furthermore, herpesvirus latency transcripts are prime targets for uncovering novel roles for non-coding RNAs, as they are often expressed in the absence of viral proteins. Projects will be available in the lab to study both the RNA-protein and RNA-RNA interactions of these transcripts using advanced proteomic and sequencing technologies.
Price Lab in the News
Selected Publications
Replication-competent adenovirus reporters utilizing endogenous viral expression architecture
O’Brien CM, Serra L, Patterson MR, Acosta RW, Yu A, Claiborne DT, Price AM. Replication-competent adenovirus reporters utilizing endogenous viral expression architecture. Journal of Virology. 2025 Sept. 10. doi: 10.1128/jvi.01146-25
Adenovirus prevents dsRNA formation by promoting efficient splicing of viral RNA
Price AM, Steinbock RT, Di C, Hayer KE, Li Y, Herrmann C, Parenti NA, Whelan JN, Weiss SR, Weitzman MD. Adenovirus prevents dsRNA formation by promoting efficient splicing of viral RNA. Nucleic Acids Res. 2022 Feb 22;50(3):1201-1220. doi: 10.1093/nar/gkab896. PMID: 34671803; PMCID: PMC8860579.
Novel viral splicing events and open reading frames revealed by long-read direct RNA sequencing of adenovirus transcripts
Price AM, Steinbock RT, Lauman R, Charman M, Hayer KE, Kumar N, Halko E, Lum KK, Wei M, Wilson AC, Garcia BA, Depledge DP, Weitzman MD. Novel viral splicing events and open reading frames revealed by long-read direct RNA sequencing of adenovirus transcripts. PLoS Pathog. 2022 Sep 12;18(9):e1010797. doi: 10.1371/journal.ppat.1010797. PMID: 36095031; PMCID: PMC9499273.
Direct RNA sequencing reveals m6A modifications on adenovirus RNA are necessary for efficient splicing
Price AM, Hayer KE, McIntyre ABR, Gokhale NS, Abebe JS, Della Fera AN, Mason CE, Horner SM, Wilson AC, Depledge DP, Weitzman MD. Direct RNA sequencing reveals m6A modifications on adenovirus RNA are necessary for efficient splicing. Nat Commun. 2020 Nov 26;11(1):6016. doi: 10.1038/s41467-020-19787-6. PMID: 33243990; PMCID: PMC7691994.
Adenovirus-mediated ubiquitination alters protein-RNA binding and aids viral RNA processing
Herrmann C, Dybas JM, Liddle JC, Price AM, Hayer KE, Lauman R, Purman CE, Charman M, Kim ET, Garcia BA, Weitzman MD. Adenovirus-mediated ubiquitination alters protein-RNA binding and aids viral RNA processing. Nat Microbiol. 2020 Oct;5(10):1217-1231. doi: 10.1038/s41564-020-0750-9. Epub 2020 Jul 13. PMID: 32661314; PMCID: PMC7529849.
Epstein-Barr virus ensures B cell survival by uniquely modulating apoptosis at early and late times after infection
Price AM, Dai J, Bazot Q, Patel L, Nikitin PA, Djavadian R, Winter PS, Salinas CA, Barry AP, Wood KC, Johannsen EC, Letai A, Allday MJ, Luftig MA. Epstein-Barr virus ensures B cell survival by uniquely modulating apoptosis at early and late times after infection. Elife. 2017 Apr 20;6:e22509. doi: 10.7554/eLife.22509. PMID: 28425914; PMCID: PMC5425254.







