Kavitha Sarma, Ph.D.
-
Associate Professor, Genome Regulation and Cell Signaling Program, Ellen and Ronald Caplan Cancer Center
Sarma studies the mechanisms of RNA-mediated epigenetic gene regulation to understand how the loss of chromatin modifier-RNA interactions impacts cellular function.
Sarma completed her graduate studies with a Ph.D. in biochemistry from Rutgers University. She conducted her postdoctoral training at the Massachusetts General Hospital-Harvard Medical School, and joined The Wistar Institute in 2016 as an Assistant Professor.
The Sarma Laboratory

The Sarma Laboratory
The Sarma laboratory is interested in the mechanisms of epigenetic gene regulation, or how the dynamic modifications of the architecture of chromatin, the complex of DNA and proteins within the nucleus of our cells, impacts gene expression and cellular function. The lab investigates consequences of epigenetic alterations in neuronal cancers and neurodegenerative diseases using a combination of biochemistry, cell and molecular biology with genome wide approaches to gain mechanistic insight into how chromatin architecture is modified in disease. The goal is to identify new pathways and interactions that can be targeted to correct these epigenetic perturbations.
-
Staff Scientist
Rahul Sureka, Ph.D.
-
Postdoctoral Fellows
Michelle Lynskey, Ph.D.
Shachin Dissanayaka Mudiyanselage, Ph.D.
Anchala Pandey, Ph.D.
Phillip Wulfridge, Ph.D. -
Graduate Students
Emanuel Forciniti
Kelvin Okpokpo -
Research Assistants
Kuo-Chen Fang
-
Lab Technicians
Afrora Muca
-
Learn about job opportunities at The Wistar Institute here.
Research
We are interested in understanding the molecular mechanisms of RNA mediated epigenetic gene regulation. Aberrations in epigenetic gene silencing can be a causal mechanism of numerous human disease and developmental syndromes. We use a combination of biochemical, cell biological and functional genomics approaches in embryonic stem cell, neural stem cell, and cancer cell models to elucidate the molecular mechanisms and functional implications of RNA containing chromatin structures in gene regulation and in genome organization.
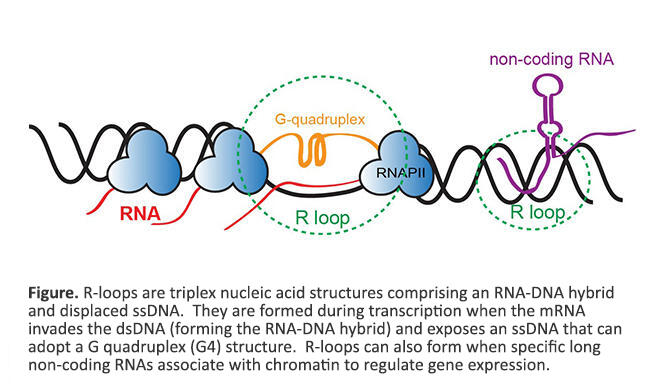
We are fascinated by triplex nucleic acid structures known as R-loops, that are comprised of a DNA:RNA hybrid and displaced ssDNA. R-loops are formed during transcription when the mRNA invades dsDNA (forming the DNA:RNA hybrid) and exposes a ssDNA that can then adopt a G quadruplex (G4) structure (see figure below). Transcription from G rich repetitive regions results in the formation of G4 DNA that impedes the reannealing of DNA strands, promotes DNA:RNA hybridization, and stabilizes R-loops. In addition to known regulatory roles, R-loops are closely linked to increased DNA damage and genome instability. Stable aberrant R-loops have also been discovered in several neurological disorders, neurodegenerative diseases, and cancers. Discovering the genome-wide locations of R-loops is challenging because of the requirement for large sample size and inefficient enrichment using the monoclonal antibody that recognizes the RNA:DNA hybrid within R-loops. We have developed a new antibody independent approach, called MapR, to identify native R-loops genome-wide. Some questions that we are interested in exploring are:
- Where do R-loops form in specific disease states?
- How do unscheduled R-loops contribute to neurodegenerative diseases and cancers?
- What are the protein factors that function in R-loop resolution and stabilization?
- How can R-loops impact gene regulation and genome organization in disease states?
- Do long non-coding RNAs localize to chromatin through R-loop formation?
Sarma Lab in the News
Selected Publications
Proximity Labeling Identifies a Repertoire of Site-specific R-loop Modulators.
Yan, Q., Wulfridge, P. Doherty, J., Fernandez-Luna, J.L., Real, P.J., Tang, H., Sarma, K. “Proximity Labeling Identifies a Repertoire of Site-specific R-loop Modulators.” Nat Commun. 2022 Jan 10;13(1):53. doi: 10.1038/s41467-021-27722-6.
A nuclease- and bisulfite-based strategy captures strand-specific R-loops genome-wide.
Wulfridge, P., Sarma, K. “A nuclease- and bisulfite-based strategy captures strand-specific R-loops genome-wide.” Elife. 2021 Feb 23;10:e65146. doi: 10.7554/eLife.65146.
Disruption of ATRX-RNA interactions uncovers roles in ATRX localization and PRC2 function.
Ren, W., Medeiros, N., Warneford-Thomson, R., Wulfridge, P., Yan, Q., Bian, Y. Sidoli, S., Garcia, B.A., Skordalakes, E., Joyce, E., et al. “Disruption of ATRX–RNA interactions uncovers roles in ATRX localization and PRC2 function.” Nat Commun. 2020 May 6;11(1):2219. doi: 10.1038/s41467-020-15902-9.
Mapping native R-loops genome-wide using a targeted nuclease approach.
Yan, Q., Shields, E., Bonasio, R., Sarma, K. “Mapping native R-loops genome-wide using a targeted nuclease approach.” Cell Rep. 2019 Oct 29;29(5):1369-1380.e5. doi: 10.1016/j.celrep.2019.09.052.
ATRX directs binding of PRC2 to Xist RNA and Polycomb targets.
Sarma, K., Cifuentes-Rojas, C., Ergun, A., Del Rosario, A., Jeon, Y., White, F., Sadreyev, R., Lee, J.T. “ATRX directs binding of PRC2 to Xist RNA and Polycomb targets.” Cell. 2014 Nov 6;159(4):869-83. doi: 10.1016/j.cell.2014.10.019.