Xiaoyu (Ariel) Zhou, Ph.D.
-
Assistant Professor, Vaccine & Immunotherapy Center
-
Genome Regulation and Cell Signaling Program, Ellen and Ronald Caplan Cancer Center
Dr. Xiaoyu Zhou is an Assistant Professor in The Wistar Institute’s Vaccine & Immunotherapy Center and Ellen and Ronald Caplan Cancer Center. She is also a Wistar Institute Assistant Professor in the Dept. of Pathology & Laboratory Medicine at the University of Pennsylvania.
She received her B.S. in Biological Science from China Agricultural University, Beijing, China, and her Ph.D. from Fudan University, Shanghai, China, where she was trained in cancer immunology. Prior to that, Dr. Zhou was a postdoctoral fellow at Yale University, where she developed immune engineering strategies and CRISPR-based multiplexed genome editing platforms to dissect and reprogram immune regulation.
Her work led to the development of novel CAR-T cell designs and genetic tools to study immune regulations in vivo. Building on this foundation, the Zhou Lab integrates immune engineering and genome editing to study intercellular immune communication and advance next-generation immunotherapies. Her research has been recognized with honors including the NIH K99/R00 Pathway to Independence Award and the Margaret Q. Landenberger Research Award.
The Zhou Laboratory
The Zhou Laboratory
The central goal of our laboratory is to understand how immune cell function is shaped by dynamic protein regulation and intercellular communication, and to harness these principles to develop next-generation immunotherapies. Our work integrates immune engineering and genome editing to interrogate and rewire immune signaling across molecular, cellular, and tissue scales.

Research
1 Immune engineering through dynamic control of immune signaling
Cell-based immunotherapies, particularly CAR-T cells, have demonstrated transformative clinical potential. However, durable responses remain limited, in part because current CAR designs do not recapitulate the dynamic regulatory mechanisms that govern endogenous immune receptors.
We previously demonstrated that CAR function can be systematically tuned by incorporating endogenous regulatory elements that control receptor dynamics. By introducing cytoplasmic regulatory motifs derived from immune checkpoint receptors into CAR constructs, we showed that receptor behavior—including antigen acquisition, fratricide, and functional persistence—could be predictably modulated, resulting in substantially improved anti-tumor efficacy in vivo (Zhou et al., Nature Immunology, 2023). This work established that engineering receptor dynamics, rather than only downstream signaling domains, represents a powerful and underexplored axis for immune cell design.
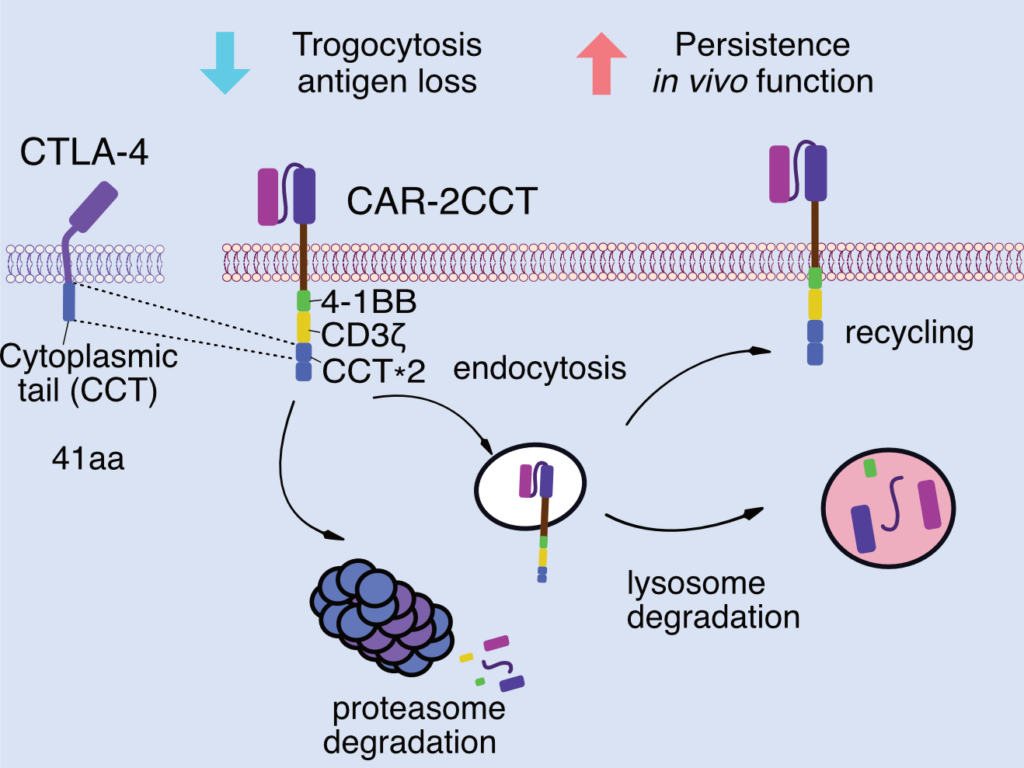
Building on these principles, ongoing work in the lab extends beyond cell-intrinsic receptor regulation to investigate how immune cell behavior is shaped by direct interactions with other cells in complex tissue environments. Rather than focusing solely on blocking suppressive pathways, we are exploring strategies to reprogram intercellular signaling itself, redirecting inhibitory or dysfunctional immune cues toward productive activation. By leveraging endogenous immune regulatory processes, this work aims to establish generalizable approaches for rewiring immune communication at the level of cell–cell interactions, with broad implications for improving immune function in solid tumors and other disease contexts.
2 Gene editing to decode intercellular immune communication
In addition to engineering with synthetic receptors/proteins, a complementary direction in which we focus is developing genome editing approaches to dissect how immune cells communicate with one another and with their tissue environment. Because intercellular immune communication is governed by coordinated activity of multiple regulators across interacting cell types, understanding these processes requires experimental systems that move beyond single-gene perturbations.
Our recent work has contributed CRISPR-based genetic technologies that enable systematic interrogation of immune regulators in physiologically relevant settings. We developed genome editing platforms that enable simultaneous perturbation of multiple immune genes within the same cell in vivo, allowing functional analysis of gene–gene interactions during immune responses (Tang et al., Nature Biomedical Engineering, 2025). These tools provide a foundation for examining immune behavior as an emergent property of interacting regulatory networks rather than isolated pathways.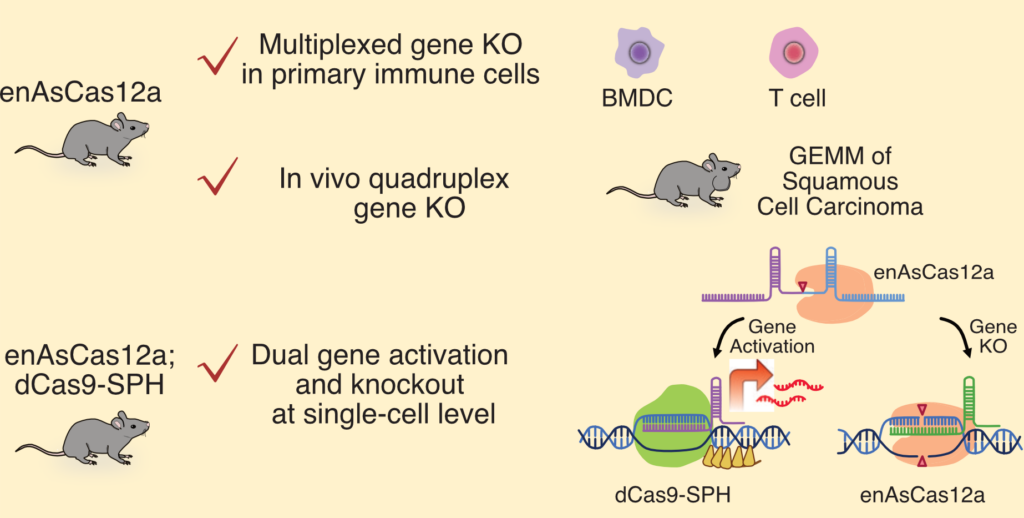
Ongoing efforts in the lab use genetic perturbation strategies to systematically probe how immune cells interpret and respond to various signals. Rather than focusing on individual pathways, this work seeks to uncover higher-order principles that govern immune communication and functional state transitions. By integrating network-level perturbation with functional measurements of immune behavior, the lab aims to identify general mechanisms that shape intercellular immune interactions and inform the development of novel immunotherapeutic strategies.
-
Research Assistant
Cole Christopher
-
Visiting Scientist
Yunfei Jiao
Selected Publications
Cas12a-knock-in mice for multiplexed genome editing, disease modelling and immune-cell engineering
Tang, K.*, Zhou, L.*, …, Zhou, X.†, Chen, S.†. High-fidelity enhanced AsCas12a knock-in mice for efficient multiplexed gene editing, disease modeling and orthogonal immunogenetics. Nature Biomedical Engineering, 2025
CTLA-4 tail fusion enhances CAR-T antitumor immunity
Zhou, X.*, Cao, H.*, Bai, M., …, & Chen, S.†. CTLA-4 tail fusion enhances CAR-T anti-tumor immunity. Nature Immunology, 2023, 24(9): 1499-1510
Applications of CRISPR technology in cellular immunotherapy
Zhou, X.*, Paul R.A.*, …, & Chen, S.†. Application of CRISPR technology in cellular immunotherapy. Immunological Reviews, 2023, 320(1): 199-216.
Systematic Immunotherapy Target Discovery Using Genome-Scale In Vivo CRISPR Screens in CD8 T Cells
Dong, M. B., Wang, G., Chow, R. D., Ye, L., Zhu, L., Dai, X., Park. J.J., Kim, H. R., Errami, Y., Guzman, C. D., Zhou, X., …, & Chen, S.†. (2019). Systematic immunotherapy target discovery using genome-scale in vivo CRISPR screens in CD8 T cells. Cell, 178(5), 1189-1204
Precise Spatiotemporal Interruption of Regulatory T-cell-Mediated CD8+ T-cell Suppression Leads to Tumor Immunity
Zhou, X., Zhao, S., He, Y ., …, & Wang, B.†. Precise Spatiotemporal Interruption of Regulatory T-cell–Mediated CD8+ T-cell Suppression Leads to Tumor Immunity[J]. Cancer research, 2019, 79(3): 585-597.
